Giant Viruses
By James L. Van Etten
The recent discovery of really, really big viruses is changing views about the nature of viruses and the history of life
The recent discovery of really, really big viruses is changing views about the nature of viruses and the history of life

DOI: 10.1511/2011.91.304
The common view of viruses, mostly true, is of tiny burglars that sneak into cells, grab the biosynthetic controls and compel the cell to make huge numbers of progeny that break out of the cell and keep the replication cycle going. Viruses are supposed to be diminutive even compared to cells that are just a micrometer (1,000 nanometers) in diameter. They are supposed to travel light, making do with just a few well-adapted genes.
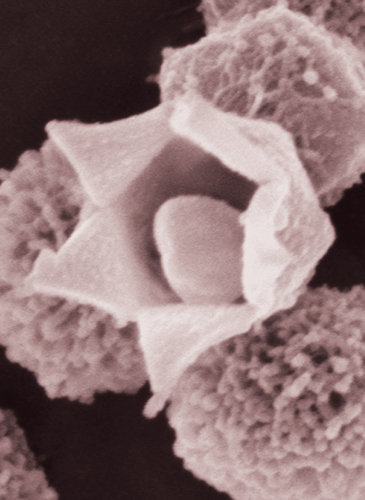
Image courtesy of Abraham Minsky, Weizmann Institute of Science.
In 1992, a new microorganism was isolated from a power-plant cooling tower in Bradford, England, where Timothy Robotham, a microbiologist at Leeds Public Health Laboratory, was seeking the causative agent of a local pneumonia outbreak. His search led to the warm waters of the cooling tower, a known reservoir for bacterial pathogens in the Legionella genus, which are the cause of the pneumonialike Legionnaire’s disease. Particles present in the sample were mistakenly identified as bacteria. Gram positive and visible under the microscope as pathogens within the particle-gobbling amoeba Acanthamoeba polyphaga, the entities surprisingly did not generate any product from the gene-amplifying polymerase chain reaction technique using universal bacterial primers.
Eleven years later, in 2003, the mystery organism received a new identity and a new name, Acanthamoeba polyphaga Mimivirus, for microbe-mimicking virus. Mimivirus is the largest virus ever discovered. Giant viruses had been known for a few years, many of them in a group termed nucleo-cytoplasmic large DNA viruses (NCLDVs). This group features several other virus families, including Poxviridae, which infects vertebrates (for example, smallpox virus) and invertebrates, the aquatic viruses Iridoviridae and Phycodnaviridae, and the vertebrate virus Asfarviridae. Giant viruses are considered to be ones with genomes larger than 300 kilobase pairs and with capsid diameters of about 200 nanometers or more.
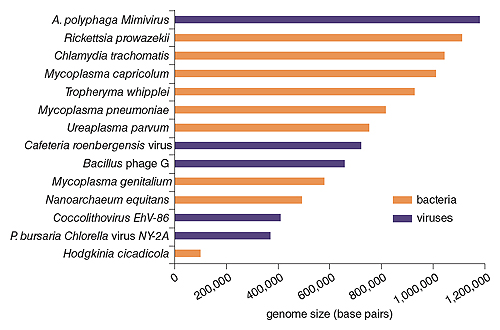
Illustration by Barbara Aulicino. Includes data provided by Alan Cann.
Mimivirus is a giant among giant viruses, with a diameter of 750 nanometers. It possesses a genome, truly outsized by viral standards, of 1.2 million base pairs, coding an outlandish 1,018 genes. For comparison, the smallest free-living bacterium, Mycoplasma genitalium, is just 450 nanometers in diameter and possesses a genome half the size of that in mimivirus, while coding just 482 proteins. The record tiniest cellular organism, Hodgkinia cicadicola, a parasite in cicadas that was described in 2009, has a genome of just 140,000 base pairs, coding a paltry 169 proteins. H. cicadicola is unable to live on its own, being entirely dependent on the lush environment of specialized cicada cells. Viruses are generally not considered living organisms (although for a consideration of their position in the phylogenetic tree of life, see the sidebar), yet mimivirus brings a bigger blueprint and more lumber to the replication process than the living H. cicadicola and many other bacteria.
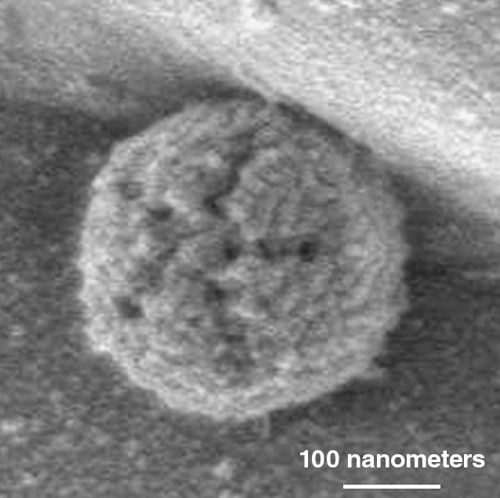

Image courtesy of Abraham Minsky, Weizmann Institute of Science.
Most giant viruses have only been discovered and characterized in the past few years. There are several reasons why these striking biological entities remained undetected for so long. Among the most consequential is that the classic tool for isolating virus particles is filtration through filters with pores of 200 nanometers. With viruses all but defined as replicating particles that occur in the filtrate of this treatment, giant viruses were undetected over generations of virology research. (Mimivirus disrupted this evasion tactic by being so large it was visible under a light microscope.) Standard plaquing procedures failed to report the presence of giant viruses because the large particles bogged down in the soft agar of the plaquing medium, disrupting diffusion and the formation of visible plaques. An additional explanation for the elusiveness of the largest viruses is that many infect protists, which have received far less research attention than plants and animals.
With the spotlight finally on them, the giant viruses are delivering striking lessons in viral physiology and ecology, not to mention challenging long-held assumptions about the shape of the phylogenetic tree of life.
Central to the placement of giant viruses in the tree of life is the presence of numerous previously unknown virus-encoded gene families. A recent reconstruction of NCLDV evolution suggests a common ancestor that likely contained a minimum set of 47 genes that have left traces in modern viral genomes. The NCLDVs then evolved by losing some genes, duplicating others and acquiring genes from their hosts and other organisms. The giant viruses, like other viruses, are genetic pickpockets, grabbing genes from their hosts over eons. Interpretation of viral phylogenetic reconstructions is therefore rife with puzzles. Yet a faded outline of evolutionary history is visible. Analysis of 45 giant viruses identified five genes common to all of the NCLDV viruses and 177 additional genes that are shared by at least two of the virus families. The ancient genetic signal, however, is very weak. Consider that in a selection of three viruses in the Phycodnaviridae family, 14 genes in common indicate a shared evolutionary history, yet within the sprawling genomes of these three biological entities, more than 1,000 total genes exist.
Mimivirus is an appropriate representative of the giant viruses, exhibiting a variety of unique properties that point to the diversity of the known giant viruses and those soon to be discovered. The mimivirus virion particle (the complete assembly of genetic material and protein coat) has an icosahedral core of ~500 nanometers covered with a ~140-nanometer layer of closely packed fibers. The fibers have not been completely characterized but based on the presence of collagen-like genes in the mimivirus genome, they may be a form of substituted collagen, the fibrous constituent of animal connective tissue. Mimivirus is the only NCLDV member known to have this peripheral fiber layer. Another singular feature of the mimivirus virion is a prominent five-fold star-shaped structure on one icosahedral vertex called the stargate portal.
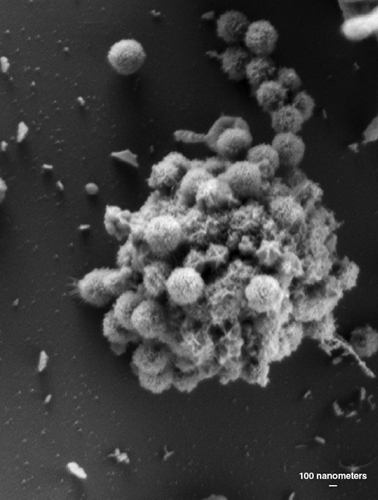
Image courtesy of Abraham Minsky, Weizmann Institute of Science.
Research suggests that mimivirus ingested by an amoeba enters the cell in a membranous compartment that fuses with lysosomes, which are digestive organelles. The activity of lysosomal enzymes is predicted to cause the stargate portal to open. An internal membrane within the mimivirus then apparently fuses with the surrounding membrane compartment, forming a conduit through which the viral genome passes into the cytoplasm of the host. A viral-assembly complex called a replication factory then forms in the cytoplasm around the viral core. The replication factory expands until it occupies a large fraction of the cell volume six hours after the initial infection.
In the replication factory, empty, partially assembled viral capsids without fibers undergo DNA packaging. Curiously, DNA packaging is reported to occur through faces of the viral capsid that are not the stargate—DNA entry into and exit from the virion apparently occur at different portals, which is very unusual for a virus.
In 2008, a new strain of mimivirus was isolated from another cooling tower, this one in Paris. With a slightly larger genome than mimivirus, the new strain was named mamavirus, and it brought with it a surprise—a parasite virus named Sputnik. Viral satellites, which are quite common, are subviral agents consisting of small amounts of nucleic acid whose replication depends on a viral genome. In this case, the viral companion may be imperfectly named, since Sputnik appears to be not merely a satellite but a legitimate parasite of its host. When present, it interferes with the infectivity of mamavirus in amoebae and appears to cause the formation of defective mamavirus virions, which is not the case for traditional satellite viruses. This unprecedented property and other features of its lifestyle have led to the proposal of a new group and new name, virophage, for viruses that parasitize giant viruses. A paper published in April 2011 reports on a new strain of mimivirus infected by a new strain of virophage. In the busy enterprise of giant virus research, news comes fast and often.
The origin of the NCLDV group is controversial. Upon discovery of mimivirus, some researchers, addressing its huge number of genes with no detectable resemblance to any cellular genes—86 percent of the total coding sequences in mimivirus—concluded that NCLDVs should be considered a fourth domain of life alongside the Archaea, Bacteria and Eukarya. It has been suggested that some NCLDV genes may have arisen from the same ancient gene pool from which sprang the prokaryotes and eukaryotes. (See the image below for a discussion of the place of viruses in the tree of life.)
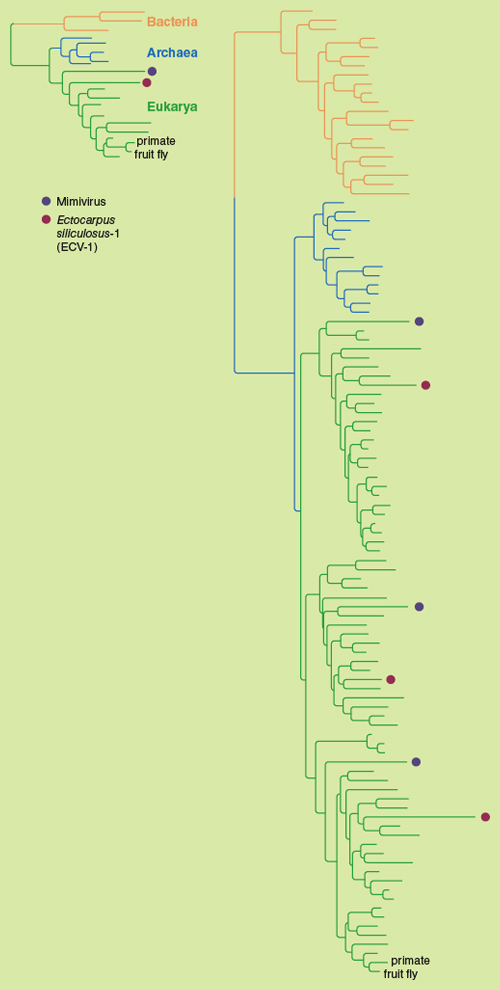
Illustration by Barbara Aulicino. Data adapted from Claverie and Ogata correspondence and from López-García and Moreira response, cited in sidebar.
An interesting pair of contrasting hypotheses suggest that given the size and complexity of NCLDV genomes, either the ancestor of modern NCLDVs may have given rise to the eukaryotic genome itself, or decay of a eukaryotic genome may have produced the genome of the NCLDV family. Horizontal gene transfer between virus and host has also played an important role in the evolution of NCLDVs (and their host cells), beginning far back in biological history.
Given their distinctness in morphology, ecology, genome size and gene uniqueness, a new name has been proposed for the giant viruses—giruses. The semantic and scientific goal of the new name is to emphasize the unique properties of large DNA viruses, which likely represent a unique and shared evolutionary history. The term virus (poison to the Greeks) is more than a hundred years old. In the time since the word was coined, a huge diversity of viral agents have been discovered with highly divergent lifestyles, physiologies and replication strategies. A collective group name worked satisfactorily when viruses were definable as small, relatively simple genetic agents dependent on hosts for their replication. The term seems less adaptable as the family of large DNA viruses is characterized in increasing detail, and evolutionary relationships between the members become increasingly visible based on deep bioinformatic analysis of their very large, complex genomes. In the past couple of years, the term girus has seen increasingly regular appearances in the virological literature.
Part of the characterization of novel biological actors is consideration of their role in shaping their environment. Giant viruses were not overlooked because they are rare, nor are they marginal players in their ecological spheres. In recent years, the emergent field of metagenomics has opened a new window on understanding ecosystems. The metagenome of an environment is the sum of all genomes of the organisms present. Using a technique called shotgun sampling, environmental material is collected, the DNA in the unsorted sample is randomly sheared, and the resulting fragments are sequenced. Overlapping sequences are then aligned to reconstruct genes. Some of the resultant genes can be identified by reference to gene databases, many cannot. The very high number of unidentified genes that are found in metagenomic studies is a driver of the surging biodiversity movement. Through metagenomics, we are in the peculiar position of knowing that the number of species we don’t know about is vast.

Loop Images/Corbis
In a striking demonstration of the power of environmental sequencing, Mya Breitbart, Forest Rohwer and colleagues demonstrated in 2002 that 200 liters of seawater contains more than 5,000 different viruses, essentially all of them unknown species. In another study, the Global Ocean Sampling Expedition sampled waters from Nova Scotia to the eastern Pacific during a two-year circumnavigation employing a more targeted approach of using known sequences of protein products, such as specific DNA polymerase fragments, to query metagenomic DNA and thus to determine the prevalence of species. In 86 percent of sample sites, mimivirus relatives were the most abundant entities after bacteriophage. Thus giruses are common in nature and it is clear that there are many more awaiting discovery.
Their role in shaping their environment is also becoming clear. More than half of all photosynthesis on Earth is carried out by photosynthetic microorganisms, including cyanobacteria and microalgae, which are collectively referred to as phytoplankton. It is estimated that at any one time, 20 percent of all phytoplankton cells are infected by viruses, including giant viruses in numbers that are unknown but clearly of quantitative importance. Central to the ecology of ocean systems, and essential to the well-being of the Earth, are microzooplankton that feed on phytoplankton and are known as protistan grazers. To date just one virus has been shown to infect a protistan grazer, Cafeteria roenbergensis virus, or CroV—a giant virus (730 kilobase pairs, 544 predicted protein-coding genes). Interestingly, CroV also has a virophage associated with it.
Phytoplankton are intimately linked to another giant virus, with consequences for sea, terrain and sky as well as phytoplankton community ecology. Emiliani huxleyi is one of the most abundant unicellular photosynthesizing algae in the oceans. Cells of E. huxleyi produce tiny scales of calcium carbonate, which, given the abundance of these microalgae, plays an important role in the carbon cycle of the ocean. E. huxleyi periodically forms huge blooms as large as 100,000 square kilometers in both the northern and southern hemispheres. A giant virus that infects E. huxleyi, called EhV (407 kilobase pairs, 472 predicted coding sequences), is largely responsible for terminating these blooms. The demise of E. huxleyi blooms releases massive amounts of organic matter, including detached calcium carbonate scales, which form large deposits. A striking example is the White Cliffs of Dover.
The termination of E. huxleyi blooms also results in the release of a chemical that is altered by other microorganisms, producing vast amounts of dimethylsulfide, which accounts for the smell we associate with sea water. When dimethylsulfide reaches the atmosphere, it induces cloud formation and rain. Thus, EhV infection of its host plays a role in the ecology, geology and climate of its environment.
The mimivirus particles in samples from the Bradford cooling tower were discovered among bacteria with the potential to cause pneumonia, and consequently there has been interest in the question of whether mimivirus might be a human pathogen.
At first glance, the odds seem unlikely that a pathogen of amoebae could make the leap to human infection. Humans and amoebae are separated by 800 million years of evolution, and an infective leap across a chasm that large would be highly unusual in virology. Typically, viral infections are highly specific for their hosts. This specificity is a result of the requirement that the virus co-opt the synthetic machinery of the host cell in order to replicate. Doing so requires many intricate and specific macromolecular interactions between viral and host components at every stage of infection, from cell entry to virus replication, which requires hijacking most of the cell’s biochemical and molecular machinery, often coupled with additional inhibition of cell defenses. It is not surprising, then, that quite similar viruses, such as HIV, the cause of AIDS, and the simian strain, SIV, do not cross-infect their closely related primate targets.
Yet mimivirus often challenges the usual rules. It gains entry into phagocytic cells (such as amoebae and possibly human macrophages) when the scavengers engulf it. It exits the vacuole that surrounds it after engulfment by relatively nonspecific membrane fusion. From that point, its huge complement of more than 1,000 genes may confer the ability to hijack or replicate cell functions beyond the ability of viruses of lesser genetic endowment.
To date, there is only slim evidence that mimivirus may infect humans. Studies in one Canadian laboratory hint that the question should remain open. Other studies find no evidence for human infection. A review in 2009 proposed that the prudent course is to treat mimivirus provisionally as a biosafety class 2 pathogen, the designation for pathogens that cause only mild disease or that are unlikely to be communicated as aerosols in a lab.
The giant NCLDV viruses probably have an ancient evolutionary history, but they are among the newest things on the scene for virologists. It should be noted that in addition to the NCLDV members, there are other viruses that qualify as giruses, including bacterial viruses such as Phage G and a virus called white spot syndrome virus, which causes a disease of major economic importance in cultured shrimp. With research efforts still in the early stages, giant viruses are already producing scientific and economics benefits. Novel enzymes are being discovered that have commercial value based on their functions and also based on the fact that viral enzymes are often the smallest in their class, making them ideal models for mechanistic and structural studies. At present, one obstacle to studying giruses is that none of them can be genetically modified by molecular techniques. Hopefully this barrier will fall soon.
Click "American Scientist" to access home page
American Scientist Comments and Discussion
To discuss our articles or comment on them, please share them and tag American Scientist on social media platforms. Here are links to our profiles on Twitter, Facebook, and LinkedIn.
If we re-share your post, we will moderate comments/discussion following our comments policy.