Imprinted and More Equal
By Randy Jirtle, Jennifer Weidman
Why silence perfectly good copies of important genes? The answer may lie in a battle between mother and father staged in the genome of their offspring
Why silence perfectly good copies of important genes? The answer may lie in a battle between mother and father staged in the genome of their offspring

DOI: 10.1511/2007.64.143
Being a diploid life form has its advantages. With two copies of each chromosome, diploid cells have a built-in insurance policy against the effects of mutation. If a gene on one chromosome has an error, there's another copy available. And for most genes, one good copy is all you need.
For this reason, geneticists have always been puzzled by the phenomenon of imprinting, in which swaths of DNA on one of a pair of chromosomes are silenced. The genes in these regions are excluded from the insurance policy. It's like flying a two-engine airplane with only one engine.

FLPA/Alamy
Gregor Mendel, the 19th-century monk who helped define genetics, never encountered imprinted genes during his studies. It's just as well he didn't, because imprinting makes a hash of his lovely laws of inheritance. Mendel was the first to explain the relation between genotype—the genes that an organism inherits—and phenotype—the traits that an organism shows. "For each character, an organism inherits two genes, one from each parent," he stated. "If the two alleles (genes) differ, then one, the dominant allele, is fully expressed in the organism's appearance; the other, the recessive allele, has no noticeable effect on the organism's appearance." These rules are part of the founding canon of genetics. But real life is often apostate, and the activity of some genes depends on which parent they came from rather than comparisons like dominant and recessive. These genes are imprinted.
At a functional level, an imprinted gene is haploid—only one allele works. It is vulnerable to the negative effects of mutations that otherwise would be recessive. Moreover, you can change its function not only with a single genetic mutation, but also with an environmentally induced change to the epigenome—the layer of heritable gene regulation not tied to DNA sequence. An epigenetic change alters the phenotype without changing the genotype. As a result of their unique genetic make-up, imprinted genes act as nodes of susceptibility for asthma, cancer, diabetes, obesity and many behavioral and developmental disorders—a list that is surprisingly long given the limited number of imprinted genes identified so far. The potential for malign influence at these sites is disproportionately large. In this respect, they're similar to the tyrannical pigs in George Orwell's Animal Farm, who famously declared, "all animals are equal, but some animals are more equal than others." The same is true for genes, and it's often the imprinted genes that are "more equal" in the formation of human diseases.
The first evidence for imprinting came more than 20 years ago from experiments with mammalian embryos that had only maternal or paternal chromosomes. Their phenotypes were strikingly different. Gynogenetic embryos (containing only maternal chromosomes) developed normally, but their extra-embryonic (placental) tissues grew poorly. The embryos died at mid-gestation. Androgenetic embryos (containing only paternal chromosomes) showed severely retarded growth, but the extra-embryonic tissue proliferated. The conclusion from these studies was that normal mammalian development depended on different genes expressed only from the maternal or paternal copy—regardless of the actual sequence of DNA. In other words, even if they possessed identical DNA sequences, the male and female genomes in mammals were not interchangeable.
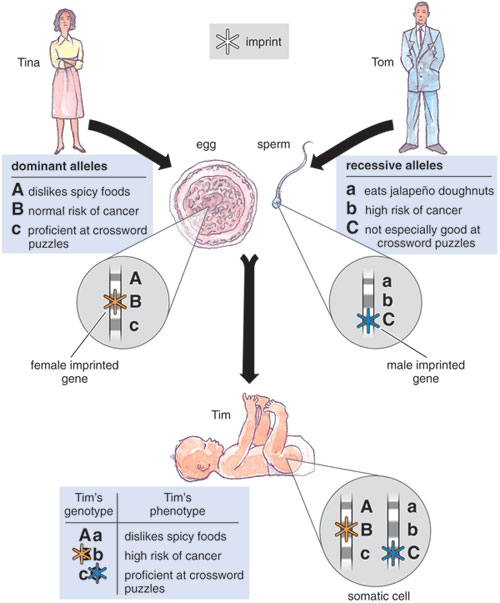
Tom Dunne
In placental, or eutherian, mammals, scientists have identified 83 imprinted genes to date. We suspect that the true number is actually much higher, as our computer-based analysis of the mouse genome predicts about 600 imprinted genes. Admittedly, no one understands fully how genomic imprinting is established and maintained at these sites. But it's clear that some kind of DNA-marking system must allow parental alleles to be distinguished from each other. Once in place, the marks in a parent's germ cells (sperm or eggs, also known as gametes) have to be maintained during fertilization and the multitude of cell divisions over the offspring's lifetime. These imprints must also be erasable and easy to reestablish during the offspring's own germ-cell formation in order to carry imprinting to the following generation.
Imprinted genes share many characteristics, including physical proximity. Most of these genes are found in clusters, an arrangement that probably reflects the nearness of regulatory DNA sequences. These clusters often contain active genes that are transcribed, or copied into an RNA format. But that RNA is not translated—not used, that is, as blueprints for proteins. Although they're untranslated, such RNA transcripts are important; without them, proper imprinting is lost. Genes for two other types of RNA molecules, so-called small nucleolar RNAs and microRNAs, are also found in parts of the genome that contain imprinted genes. Although their exact roles remain mysterious, scientists speculate that these RNAs help control the activities of imprinted genes by preventing the target RNA from being translated into protein.
Epigenetic factors also help establish and maintain genomic imprinting by controlling how tightly the chromatin—the combination of DNA and protein that makes up a chromosome—is coiled. Tightly coiled, or condensed, chromatin restricts gene activity, whereas a more open configuration creates a more permissive environment for genes to be turned on. There are several molecular tools used to regulate chromosome condensation. Methylationentails the covalent attachment of methyl groups to DNA, whereas phosphorylation and acetylationstick other small molecules onto histone proteins, the spools around which DNA is wound. Methylation and phosphorylation restrict access to genes by winding the DNA and histones tighter; acetylation does just the opposite. In addition to these chemical modifications, a handful of non-histone proteins also bind to DNA and regulate gene activity.
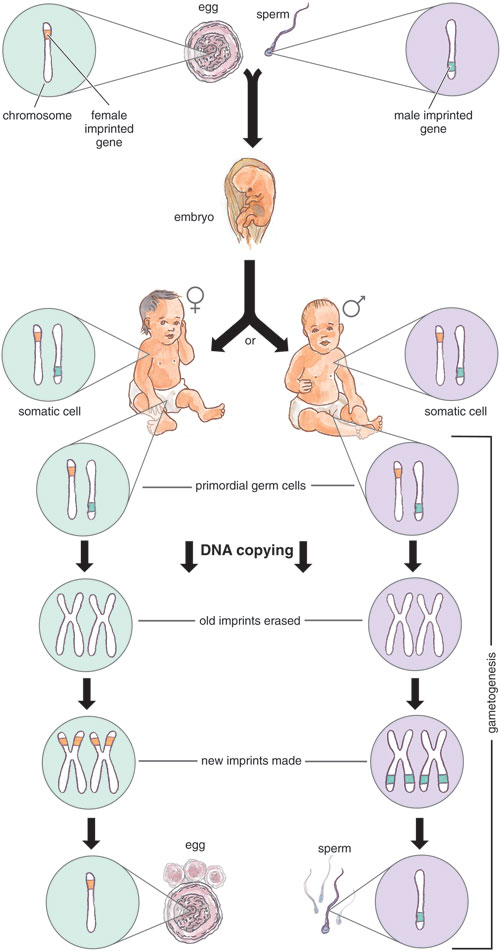
Tom Dunne
Many of the epigenetic changes that regulate imprinting take place in or around so-called CpG islands, regions of DNA that have many cytosine-guanine, or CG, base pairs. (The DNA molecule has a phosphate group between adjacent nucleotide bases, hence the "p.") The methylation state of CpG islands in eutherian genomes can vary from complete, as it is near the DNA left over from old virus attacks, to nonexistent, as it is near the genes used most frequently. In between these extremes are CpG islands that are sometimes methylated and sometimes not. Some of these so-called differentially methylated regions control imprinting.
The methylation reaction itself is catalyzed by an enzyme called DNA methyltransferase, which connects a methyl group (CH3 -) to a cytosine in DNA. This chemical change alters slightly the shape of the double helix, preventing the binding of many types of accessory proteins. Once instituted, methylation can be maintained even if the DNA is copied. In this way the genome retains its methylation pattern throughout development. In some cases it can even persist from one generation to the next.
Because the pattern of DNA methylation is both stable and heritable, many geneticists have concluded that methylation is the epigenetic basis for imprinting. Good evidence supports this connection. So-called imprint control regions near many imprinted clusters are methylated differently depending on which parent they came from. Yet finding them has proved to be a challenge. These control regions don't share a common DNA sequence, unfortunately, although they seem to be correlated with areas where cytosine-guanine dinucleotides are more frequent. Simple repeated sequences in the DNA code are often nearby, but the significance of this is unknown. Almost all imprinted regions contain differentially methylated stretches, and several experiments show that deleting these segments prevents normal imprinting.
Methylation of DNA does not, however, tell the whole story. Histone proteins also contribute. For several genes, the state of the histones in a particular region of DNA is tied to which parent that chromosome came from. Some scientists suggest that DNA methylation is connected mechanically to histone modification. Methylation of CpG islands can recruit other proteins that bind the DNA and, in turn, attract still other protein enzymes that remove the acetyl groups from histones. This complex of proteins condenses chromatin and limits transcription. Evidence keeps accumulating that histone modifications help to distinguish parental alleles. But most scientists still consider DNA methylation the primary means of maintaining the epigenetic memory of imprinting.
Despite the genetic vulnerability that imprinting dictates, every placental mammal studied so far has retained this attribute in its genome. Clearly there must be some advantage that offsets the risk, although exactly what this benefit might be has generated much philosophical debate. Scientists still disagree about why imprinting evolved in the first place and what selective pressures have maintained it throughout the mammalian family tree.
One theory is that imprinting is a solution to the problem of unfertilized eggs that develop into new individuals, a phenomenon called parthenogenesis. The recent account of a captive Komodo dragon's "virgin birth" in the United Kingdom is an example of parthenogenesis, which is possible, if rare, among reptiles, amphibians and fish. It occurs more commonly among certain invertebrates and plants. According to the anti-parthenogenesis idea, the risk from imprinting a few genes is negligible compared to the genetic benefits of sexual reproduction for long-term evolutionary fitness.
A second hypothesis is that imprinting evolved as a way to defend the genome against parasitic foreign DNA. In this view, imprinted genes are the equivalent of civilian casualties, innocent bystanders that were inactivated because they were too close to a sequence that resembled a DNA parasite spliced into the genome. Yet another conjecture, the ovarian time-bomb hypothesis, argues that imprinting protects females against germ-cell tumors by guarding against excess placental growth. Each of these theories predicts that imprinting is adaptive—that it serves a specific function to help a species to survive.
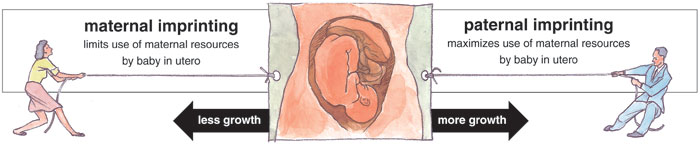
Tom Dunne
However, most scientists do not think imprinting is a beneficial adaptation. The most widely accepted theory of its origin, called the conflict hypothesis, argues that imprinting is the unintended result of a reproductive battle between the sexes. The important elements in this war are polyandry (females mating with more than one male in a breeding season), live birth and the much greater investment that mammalian mothers make in their offspring, compared to mammalian fathers. According to this provocative idea, imprinting arose because of a genetic tug-of-war over the amount of nutrients extracted from the mother by her offspring. The conflict hypothesis predicts that genes that are only active on paternal chromosomes will promote prenatal growth, thereby maximizing the evolutionary fitness of the offspring. By contrast, genes that are only active when inherited from the mother suppress offspring growth in order to maximize the mother's reproductive success—which, by definition, involves more than just one offspring.
It is important to note that these theories are about the evolutionary logic of imprinting rather than the molecular mechanisms themselves. In fact, none of the current models provides a good mechanistic framework for imprinting, a flaw that weakens their predictive power. Reconstructing the origin of genomic imprinting will, in the end, require an analysis that combines the evolutionary framework and the possible mechanisms.
Coincidentally, the first imprinted genes to be identified provide some of the best support for the conflict hypothesis. Insulin-like growth factor 2, or IGF2, and its receptor, IGF2R, direct cell growth and development throughout an organism's life. We discovered that neither of the genes that encode these proteins are imprinted in birds or in monotremes—mammals that lay eggs, such as the platypus and echidna. Yet both genes are imprinted in the remaining mammals (therians), which do not lay eggs. Thus, imprinting at these two sites in the genome must have evolved with the development of live birth roughly 180 million years ago.
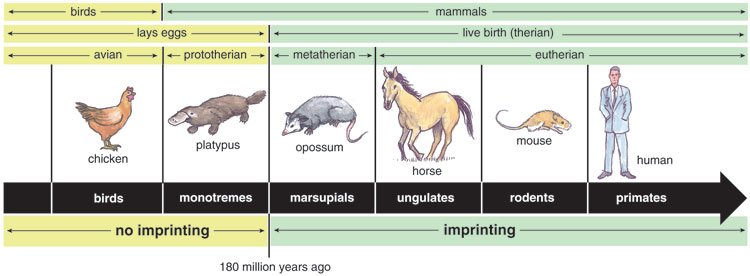
Tom Dunne
Looking at the imprinted regions around the IGF2 and IGF2R genes in mammals from across the evolutionary spectrum has enabled us to determine how these physical changes to DNA first evolved and how the changes are made and maintained today. In the past, this kind of comparative analysis has had limited success in identifying imprint-specific sequences because the mammalian species used in the analysis were too closely related. We looked at the genomes of distant relatives— opossum and platypus in addition to human and mouse. The genomic similarity we found at the site that regulates imprinting of the IGF2 gene indicates that the mechanism of imprinting has not changed much since the first stages of therian radiation.
Nevertheless, differences do exist. One of our most surprising discoveries was that in marsupials such as the opossum, most of the control elements identified for IGF2R imprinting in eutherian mammals are missing—yet this gene continues to be imprinted. Thus, either there is an ancestral mechanism of IGF2R imprinting in both marsupials and eutherians that has not yet been identified, or independent mechanisms evolved to imprint this gene in these two mammalian groups. This fundamentally important conclusion suggests that the regulation of imprinting is more complex than that we currently understand.
Imprinting patterns diverge at other genomic sites too. For example, the gene delta-like 1 homolog (DLK1) is imprinted in eutherian mammals but not in marsupials. And the neuronatin gene (NNAT), which is also imprinted in eutherians, is not even found among non-eutherian mammals. Imprinting isn't a one-way street, either. The human gene for IGF2R is no longer produced from one allele as it is in most other mammals. Judging by the absence of IGF2R imprinting among humans, tree shrews and flying lemurs, the imprinting of IGF2R disappeared about 75 million years ago in one of our common ancestors. Thus, IGF2R is imprinted in mice but not in humans. These observations suggest that being able to change the imprint status of genes and make epigenetic changes to the genome may have a critical role in speciation.
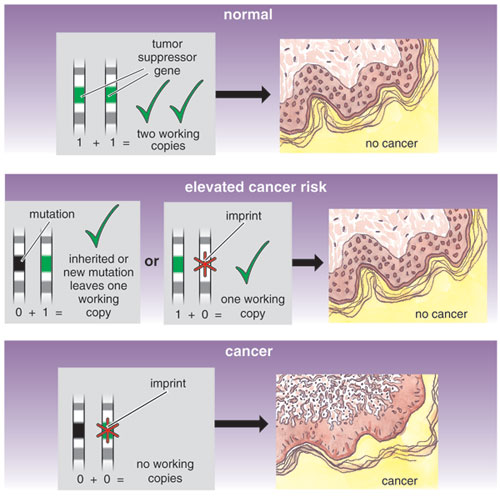
Imprinted regions are effectively haploid, which makes them vulnerable to recessive mutations and epigenetic changes. Many developmental disorders and human diseases are linked to imprinted genes. Furthermore, because these genes are often physically close to one another and controlled together, a single epigenetic or genetic change in the region can disrupt many genes. Environmental dysregulation during early development can also affect imprinting-control elements, resulting in chronic diseases that persist into adulthood. For example, Beckwith-Wiedemann syndrome, which is characterized by organ overgrowth, can be caused by mutations in any of several imprinted genes clustered together on part of the short arm of chromosome 11. But the same syndrome can also arise from epigenetic changes, such as imprinting patterns, that alter the activities of genes within the cluster—despite the fact that the DNA sequence itself is unchanged.
Tom Dunne
In another classic example of imprinting-related disease, the same mutation can yield either Prader-Willi or Angelman syndrome, two serious but very different disorders, depending on whether the altered gene comes from the father or mother. When it is paternally inherited, a mutation to one specific gene in an imprinted region on chromosome 15 gives rise to Prader-Willi syndrome. Maternally inherited mutations in the same gene—while themselves silent owing to the imprint—lead to improper repression of a nearby gene that is also imprinted, causing Angelman syndrome. Sadly, it appears that the incidence of imprinting-related developmental disorders is significantly increased in children conceived by in vitro fertilization, a pattern that points to the delicacy of sustaining genomic imprints during gamete fusion and the first few cell divisions.
Tom Dunne
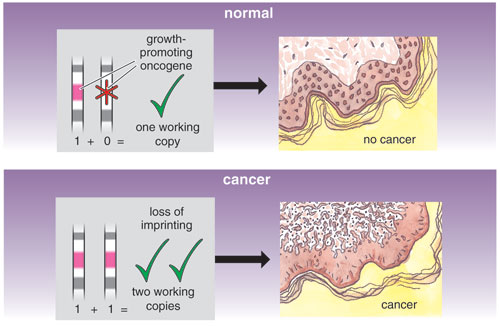
Imprinted genes are often involved in cancer pathogenesis as well. A silent, imprinted allele is often equated with the first hit in geneticist Alfred G. Knudson's now-famous "two-hit hypothesis" of cancer development. This theory states that since most of the genes that regulate cancer are still effective even when one of the two copies is disabled, going from a normal phenotype to a cancer phenotype actually requires two mutations, or "hits" to a cancer-controlling gene. A preexisting mutation in one of the genes would act as one hit; so would an imprint. Indeed, aberrant activity among imprinted genes is linked to a range of carcinomas, including hepatoblastomas, rhabdomyosarcoma and adrenal carcinoma. For many patients with Beckwith-Wiedemann syndrome, abnormal methylation of one gene on chromosome 11 correlates with the development of pediatric cancers. And several adult-onset cancers, including colorectal carcinoma, bladder cancer, osteosarcoma, ovarian cancer and breast cancer, are linked to IGF2 overproduction because of the loss of imprinting.
Some complex behavioral traits also seem to have an imprinted component. An example comes from females with Turner syndrome, who have only a single X chromosome that can come from either their mother or father. Depending on the origin of their X chromosome, they may develop atypical cognitive and social phenotypes. This inheritance pattern points to the presence of one or more paternally activated genes on the X that regulate these dimensions of personality. And because normal XY males only inherit the X chromosome from their mother, the presence of such a maternally silenced allele could explain in part the sexual dimorphism in sociability between males and females. Parent-of-origin effects also show up in other neurobehavioral conditions, including autism, Alzheimer's disease, bipolar disorder and schizophrenia. These findings have led to the intriguing hypothesis that such disorders stem from imprinting errors during early brain development. The imprinted genes involved in the formation of these disorders are unknown. Nonetheless, it is apparent that many imprinted genes in the human genome remain to be discovered.
Until geneticists understand much more about the molecular evolution of imprinting, the debate about whether imprinted genes are adaptive or maladaptive will continue. Yet both camps would agree that this subset of genes needs to be identified in humans because of its often-harmful influence. Toward this end, we are now applying to the human genome the same algorithms we originally developed to find imprinted genes in the mouse genome. By mapping those genes predicted to be imprinted in humans onto the landscape of disease risk defined by linkage studies, we hope to define the genetic—and epigenetic—components of many more diseases.
Click "American Scientist" to access home page
American Scientist Comments and Discussion
To discuss our articles or comment on them, please share them and tag American Scientist on social media platforms. Here are links to our profiles on Twitter, Facebook, and LinkedIn.
If we re-share your post, we will moderate comments/discussion following our comments policy.